My last couple of weeks have been full of sharing and swapping techniques and tips with new and visiting members of the Perry lab, Anth and Biology departments. It's ranged from advice for field work in Panama to how we in the Perry lab prefer certain brands of micropipette tips over others. It's got me thinking about ways I like sci comm-ing with other researchers, which will be the theme of this week's blog post.
| When starting work at a new job and/or lab, your first tasks typically revolve around learning "how we do things here". I've been managing the Perry lab modern space for a little over a year, and it shows a bit (as you'll see on the right). I love labels and standard systems for organization. It makes training and acclimating researchers that much easier if they are confident they'll be able to find everything on their own - no one enjoys feeling like they're bugging you with silly questions. I also am a big fan of checklists (see below). Ticking boxes and crossing things out makes me feel like I'm accomplishing stuff, even if it's little steps in a protocol. |
For any regularly occurring protocol, like shearing DNA fragments to smaller sizes using a machine in a different building (below left) or TruSeq Nano DNA Library Preparation (below right), I'll either make my own checklist or take detailed notes on a pre-existing protocol, then put the prints into one of these clear plastic binder sleeves. That way, I can use a dry-erase marker to cross off steps or check things off the list over and over again without wasting paper. Go green or go home!
| One of the benefits of reusing protocols in a binder such as mine is that it makes your lab notebook a lot less cluttered. For those who are unfamiliar, researchers almost always have a logbook that stays in the lab space in which they record exactly what they do/observe/change. Such a practice is meant to reinforce the idea of repeatability - if the results of an experiment are unusual or controversial or particularly interesting for whatever reason, another researcher could repeat exactly what you did to see if they achieve the same results. If you use "laminated" protocols like me, you could keep said protocol with your notebook and then only have to worry about recording sample IDs, alterations you made, and results :D |
First piece of advice I give to anyone new in the lab is to get a notebook that suits their needs. I prefer these because I like gridlines and easy organization, but these are nice if carbon-copy pages will ever be useful to you (ie, keeping a record for yourself outside the lab - super useful for our ancient DNA lab space). Don't be afraid to tape things in either! Just don't forget to keep a digital record of everything for easy sharing with collaborators.
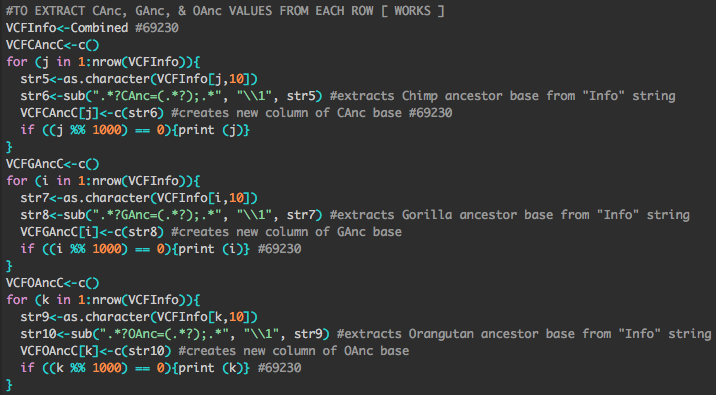
| Speaking of digital records, I am a big fan of annotating any bioinformatics codes I write. That said, the result is almost always something that makes sense to me but might not be the most accessible to others. There's a slim chance you'd know what all that gray text was talking about on the right, but it's not easily kept in check. |
More effective and brilliant coders, like Perry lab post-doc Christina Bergey, prefer systems like GitHub for controlling various versions of their code. Don't worry, I've made a profile and will now work towards bettering myself in the world of writing code for things. First step there is to become a better bioinformatician. Time to learn Python...
How do you like to organize your work? Any suggestions always welcome :) Cheers!




 RSS Feed
RSS Feed
