Hi friends! I'd like to start this blog by thanking you for visiting me/my website again after my 2-month lag on posts. To all newcomers (probably from #Evol2019), hello and welcome! Please know that I appreciate all of you :)
If you are visiting my blog from Evol2019 and seeking more information about my poster, feel free to skip my usual blog nonsense and scroll down a bit!
| I'm a little behind on my weekly posting schedule for two reasons. First is personal; my family members were getting cut open and put back together again all within the span of a few weeks. At the end of March my little fluff butt had a blocked urethra and needing emergency veterinary care for a few days. Then in the second week of April my Momma had her hip replaced, and I went back to NJ to help her with her recovery. I'll write up some tips for both of these situations over in the Life Updates section soon. |
Second is research-related; I had a conference coming up and needed to get a wiggle on in my lizard dissertation work. Some of y'all might remember my blog about Evolution - Montpellier 2018 - we're gonna make a few adjustments to my Conference 5 Ws:
| WHAT |
| WHERE |
| WHEN | 21-25 June 2019 (Yup, I'm blogging from the conference! Happy to meet up with you for a coffee or something if you're here too!) |
| WHY | To partially quote myself from the Evol2018 blog, "Because evolutionary biology is awesome and a lot of people wanted to talk about it all at once in a pretty awesome town in" New England :) |
I love this conference for so many reasons, one being the sheer variety of topics that are covered. The first day alone I heard about cascading community consequences of fish adaptation [Talk693], the health of ancient humans [Talk229], the stability of white shark genomes [Talk403], and inheritance of stripy color patterns in skinks [Talk226], just to name a few. I'm gonna quote my Evol2018 blog again: I highly encourage you to check out the #Evol2019 Twitter hashtag so you can see some of the materials that were presented. A fair portion of the talks were even recorded; I'll link to those once they are available. But at the very least, you can scroll down a bit to see my poster :)
I took advantage of Jakub Nowosad's collection of colorblind friendly palettes in my efforts to make a pretty poster for this year's conference. My accent colors for each organism were themed: fiery for the fire ants and a blue I borrowed from the belly of a male fence lizard with Andrea Cirillo's paletter package. I also used some silhouettes from Phylopic, as well as some images y'all might recognize from my review paper :)
I try to write my blog posts for more general audiences, so I am just going to highlight brief key takeaway messages from each section of my poster. Please feel free to message me if you have any questions!
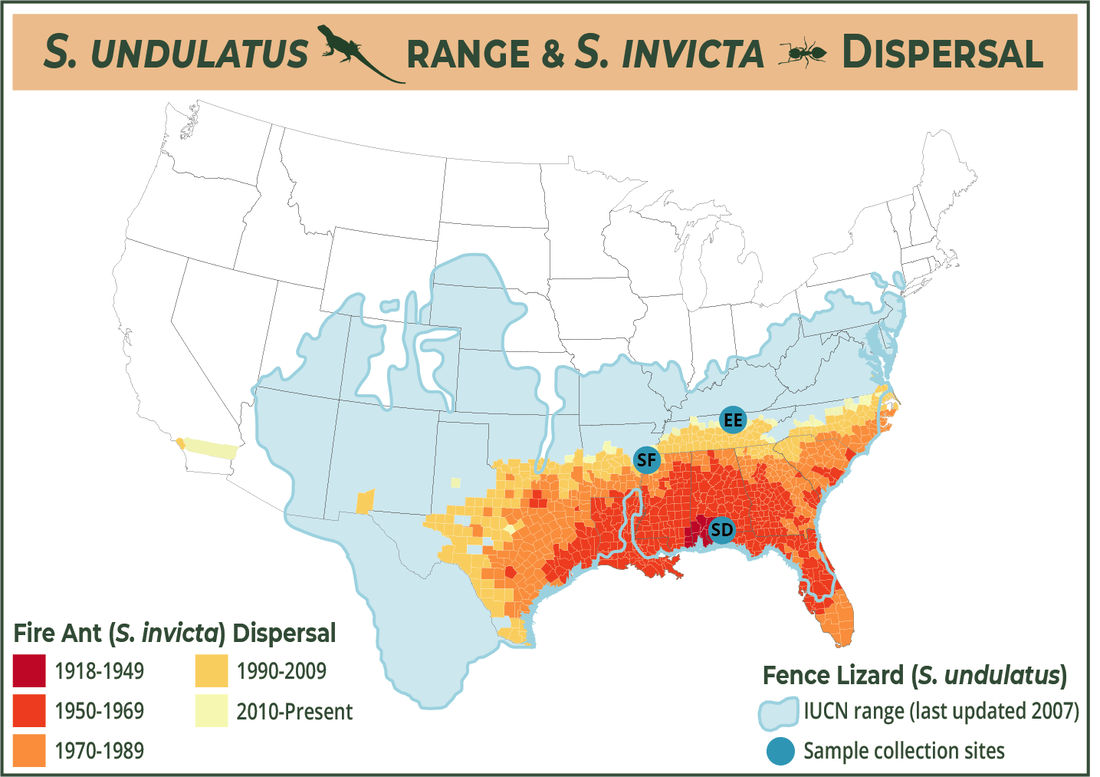
The goal of this project is to investigate some fence lizard phenotypes (AKA observable characteristics: longer hind limbs and twitchy behavior) that are associated with longer exposure time to human-introduced, invasive fire ants. Both of these phenotypes enable the lizards to escape ant predation when compared with less twitchy, shorter-limbed lizards. We want to better understand the genetic components underlying these physical changes.
I am working with lizards that Tracy Langkilde's lab has collected over the past decade from the sites shown in the map below. For our preliminary work (and my dissertation chapter) we decided to compare the genomes of a population of lizards that has been exposed to fire ants since the 1920s (that site's called Solon Dixon [SD]) to two populations of lizards with minimal fire ant exposure (Edgar Evins [EE], St. Francis [SF]). We hope to increase this work in the future to include lizards Tracy's lab has collected from other sites in the American southeast.
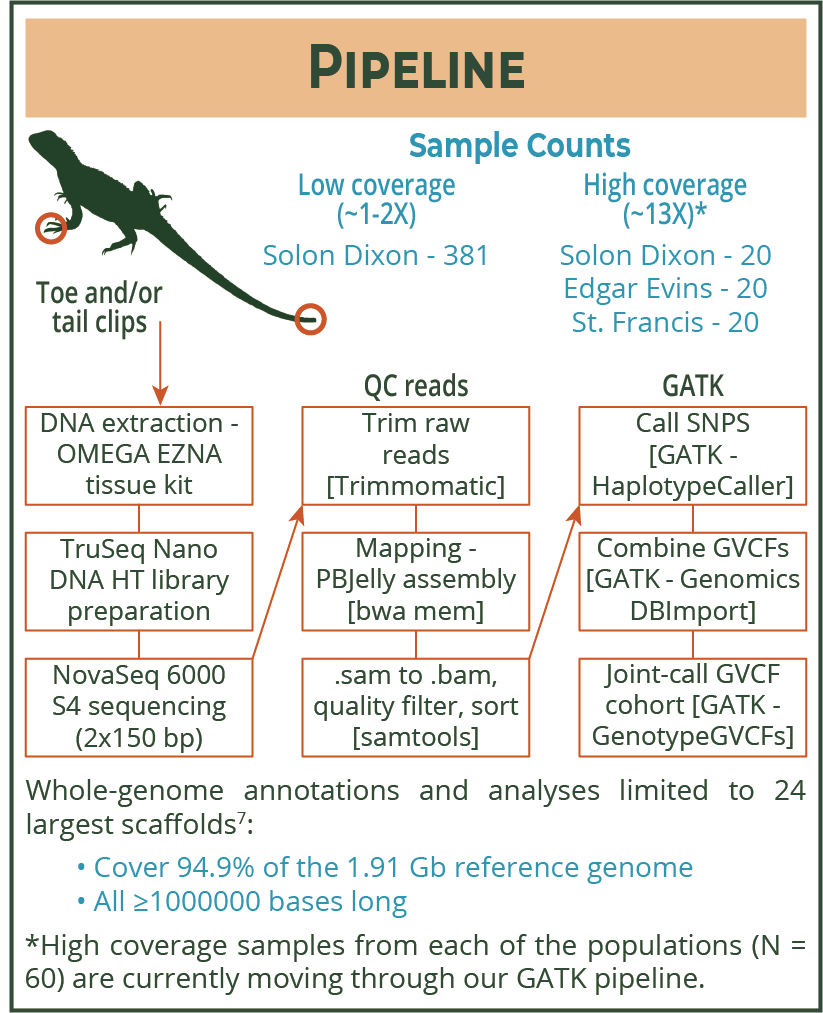
| We are working with a group of individuals led by Tonia Schwartz to publish a reference genome for this particular species of lizard. Since we had access to this genome, we decided to sequence lots of SD individuals to lower coverage - this just means that the sequencing machine read over each individual's genome only a couple of times. I could then map these low-coverage genomes to the reference genome to line up the reads that the sequencer generated. Once the reads are aligned and in the same order, I used a package called GATK to call SNPs and genotype the entire population together. If you are unfamiliar with SNPs, see this blog. |
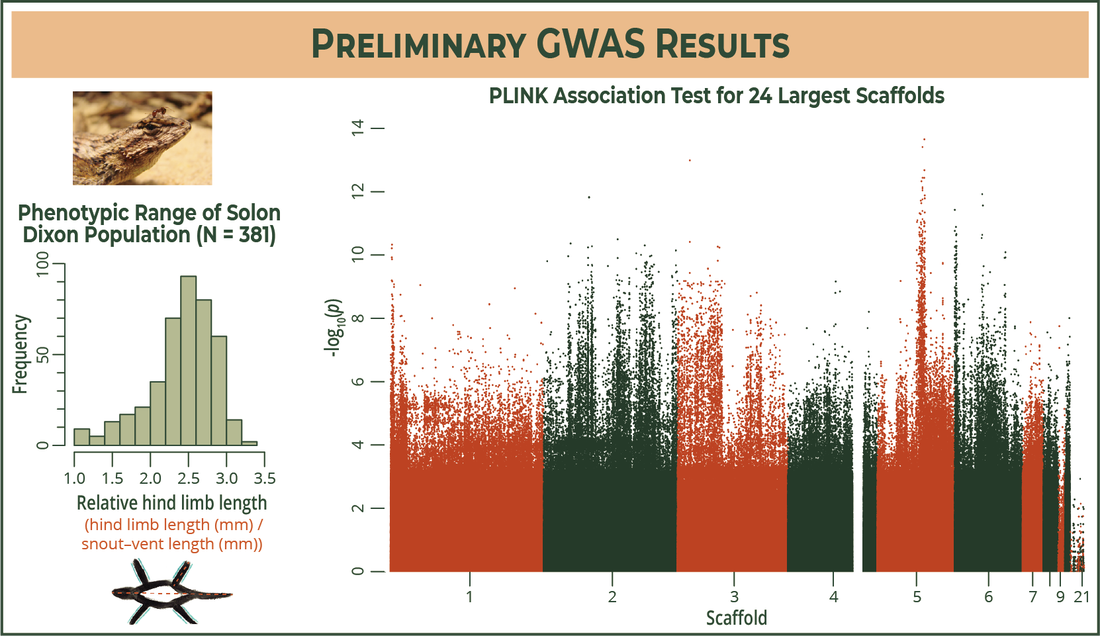
I limited my analysis to the limb-length phenotype for the poster. I used PLINK to read through the genome-wide SNPs I found and associate them with the relative hind limb lengths present in the SD population [GWAS]. Each dot on this plot is one SNP in its respective genomic location (x-axis), and SNPs with a higher association for longer limb length will have a smaller p-value, which means they'll be higher on the y-axis. I'm now looking into these peaks to see if the SNPs are in or near genes that code for things that might affect an individual's limb length, think skeletal/muscle growth and things like that.
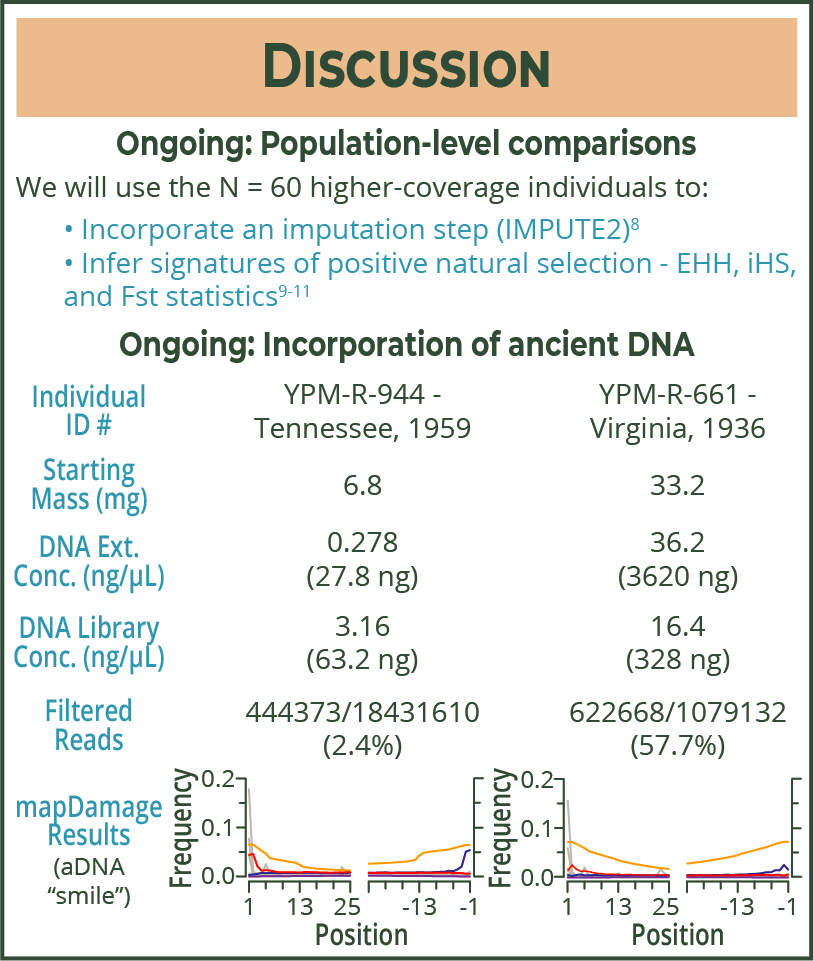
| My higher-coverage genome data is chugging along through my pipeline now. A lot of biological computational packages codes were written with really well-studied lifeforms in mind (ie, humans or model organisms like mice or fruit flies). This means that trying to use these packages for non-human, non-model organisms can be messier and/or take a lot longer. I've also submitted applications to acquire more museum samples (cuz ancient DNA in fun and exciting, and could allow us to see how allele frequencies might have changed over time as the SD lizards adapted to fire ant presence). |
| The QR box above links to a high-quality PDF of my poster. Please send me any questions you might have, happy to chat :) |
And please, if you are a current Evol2019 attendee, send me a message if you'd like to meet up on Monday or Tuesday! To all my readers, conference-goer or not, have a wonderful day: take breaks, enjoy your locale, and immerse yourself in the science that these awesome people would like to share with you. Cheers!





 RSS Feed
RSS Feed
