| This post has been a while coming, but it's here at last! After going through the processes so beautifully illustrated in this gif, my first 1st-author research article has officially been published :D An evolutionary medicine perspective on Neandertal extinction is available online through the Journal of Human Evolution (yay for another addition to my "Publications" page :D ). |
As I've mentioned on my "Research" page, I began my work on this project during my first few months as an official member of PJ's lab. I had come into the lab with absolutely zero bioinformatics experience, and what better way to learn than to try a little mini project of my own? This analysis was based on the work (and genomes) Castellano et al published in their 2014 PNAS paper Patterns of coding variation in the complete exomes of three Neandertals. I've decided to use this blog post to provide a bit of background genetics information that you'd probably appreciate if you're unfamiliar with the field but would still like to take a stab at reading the paper just because you like me :)
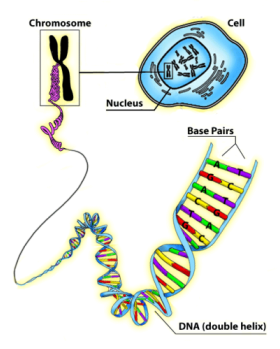
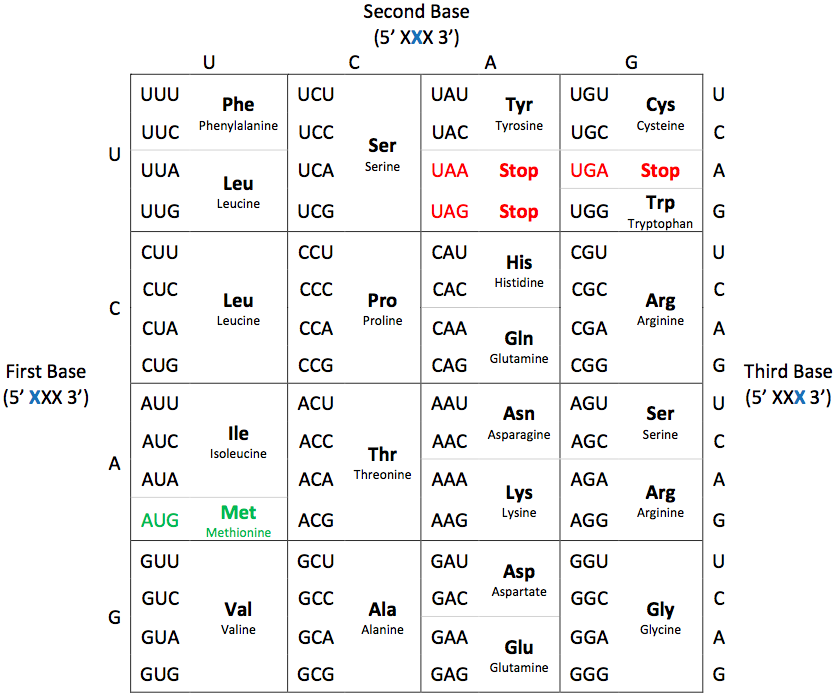
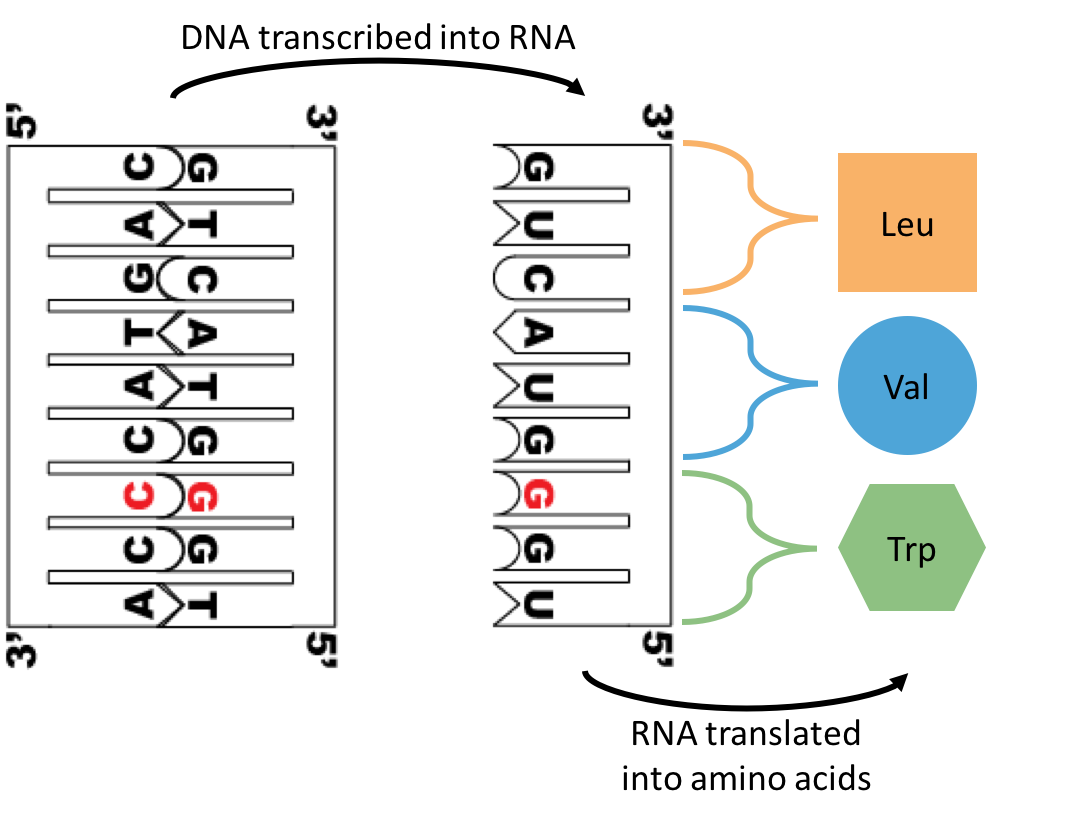
Our genetic information is carried in our DNA [deoxyribonucleic acid], which is composed of the nucleotides Adenine (A), Cytosine (C), Guanine (G), and Thymine (T) [see image above left]. During the interphase portion of the cell cycle (ie, when cell growth and development takes place), our genetic information is transcribed into RNA [ribonucleic acid]. RNA is composed of the same nucleotides as DNA, except a Uracil (U) takes the place of the thymines. Then this RNA is translated into the amino acid sequence of a protein. Proteins are made up of hundreds of amino acids, and there are only 20 amino acids to choose from [see image above right]. Here's a neat video that draws out the process for you.
| Proteins are often called the "workhorses of the cell" because they are largely responsible for the structure, function, and regulation of our tissues and organs. The motor protein featured on the left is an excellent example of a protein at work in our cells. It is therefore essential that transcription and translation correctly assemble these proteins, otherwise our bodies won't work the way they should. |
recap: DNA --> RNA --> Amino Acids --> Proteins --> US
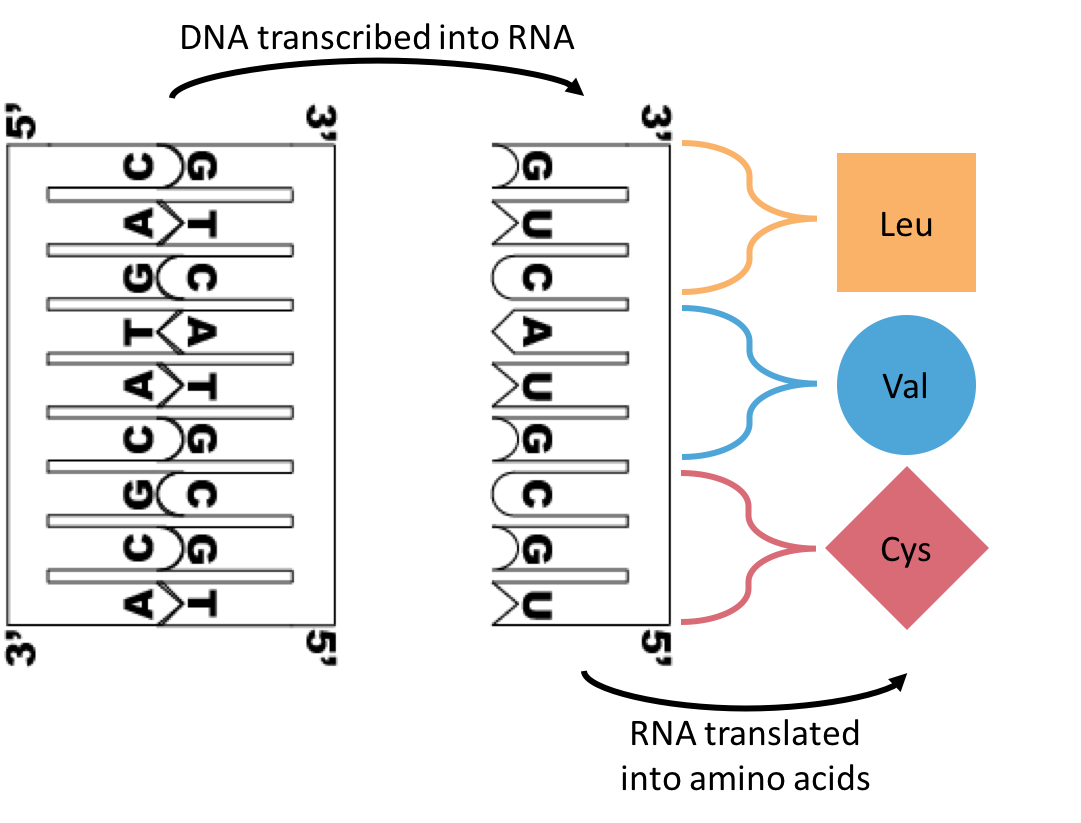
I drew out a hypothetical example of a segment of DNA going through transcription and translation to end up as three amino acids: Cysteine, Valine, and Leucine. In certain individual spots of the human nuclear genome, there are these things called SNPs [Single Nucleotide Polymorphisms]. As the name kind of suggests, a single nucleotide [A, C, G, T] can take multiple forms within the individuals of a population.
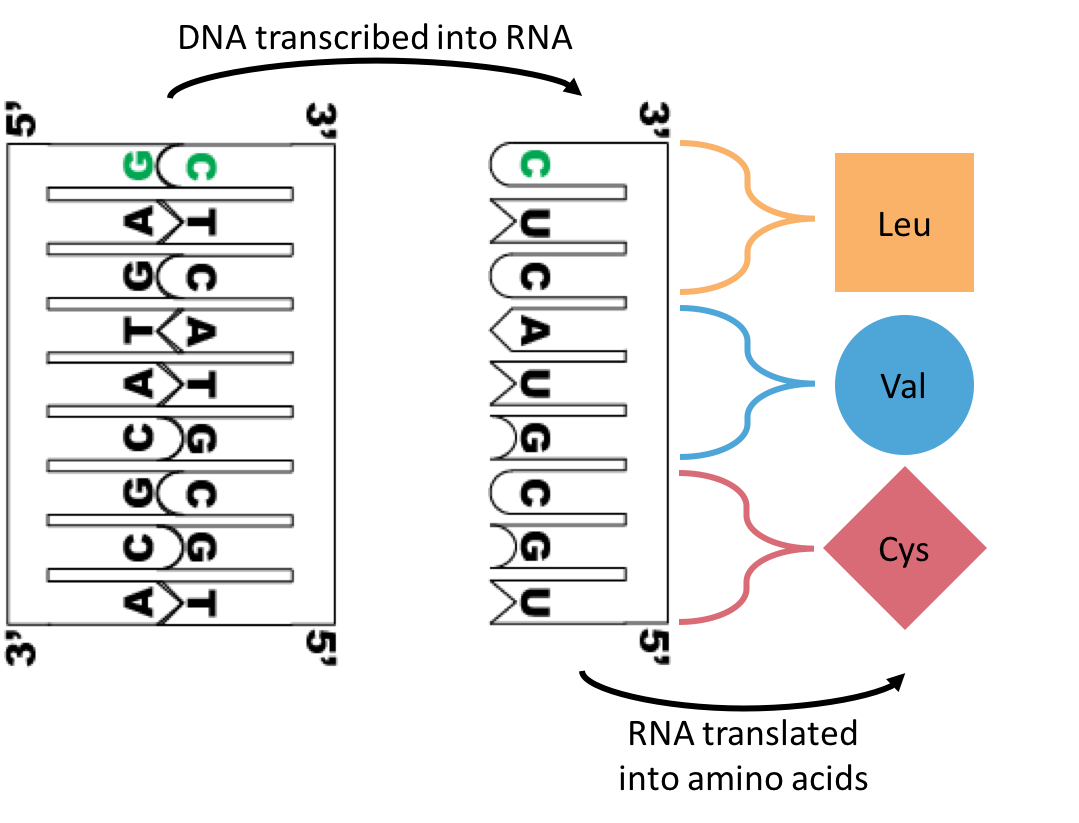
| So let's say your genome has the above DNA sequence, and my genome has the one pictured here. At the green highlighted spot, my DNA will transcribe a cytosine into my RNA where yours has a guanine. This change doesn't actually change the protein that is formed when my RNA is translated, so our proteins will probably work the same way. This is called a synonymous SNP. |
| As you might have guessed, sometimes these SNPs might cause our RNA to be translated into a different amino acid. This is called a nonsynonymous SNP, and is depicted on the left. If we have genomic data available for a population of organisms (in my case, modern humans and Neandertals), we can use a program called PolyPhen-2 to predict whether or not these nonsynonymous SNPs will functionally change ("damage") the structure of the protein. |
Castellano's group generated a dataset in which they compared the number of potentially damaging nonsynonymous SNPs in Neandertal genomes to the number found in modern human genomes. They published evidence that Neandertals had much lower genetic diversity than modern human populations from Africa, Europe, and Asia, and that Neandertals had a nearly 50:50 ratio of damaging to non-damaging nonsynonmous SNPs (a much higher proportion when compared to the modern human populations). My analysis replicated the one conducted by Castellano et al, and went a bit further by assessing these same kinds of patterns within certain gene families that are associated with innate immunity.
I gave a talk about this project at AAPA 2016 and have provided the slides here for your perusing pleasure. Otherwise check out the paper to find out more, especially since we've updated a bunch of things since we uploaded our initial bioRxiv preprint :) Enjoy!


 RSS Feed
RSS Feed
